- Home
- News
- Spotlight on Science
- Visualising breathing...
Visualising breathing motions of protein structures
07-03-2022
Proteins are not rigid structures: they move, breathe and adapt to their environment in order to optimise their shapes and functions. Cutting-edge work in NMR spectroscopy and at the ESRF's X-ray crystallography beamlines maps the structural details of “breathing motions” in the core of a protein.
Nuclear magnetic resonance (NMR) studies carried out in the 1970s surprisingly demonstrated that aromatic amino acids in proteins can undergo so-called ring flipping (i.e., 180° rotations of the aromatic side chain). Paradoxically, these aromatic amino acids are often located in the tightly packed protein core, where they engage in multiple interactions to maintain the protein fold and thereby function. At that time, it was proposed that large-scale protein “breathing motions” would be necessary to accommodate these ring flips; however, until now, the structural details of these motions have remained enigmatic.
By combining NMR spectroscopy and X-ray crystallography, researchers from the Institut de Biologie Structurale (IBS) and the Institute for Advanced Biosciences (IAB) in Grenoble have been able to map the structural changes associated with aromatic ring flipping in the core of a protein (Figure 1). Using state-of-the-art macromolecular crystallography beamlines at the ESRF (ID30A, ID23-1 and ID23-2) and at the Diamond Light Source (I04 and I04-1), the study shows how specific structural changes in the protein generate a void volume around the aromatic ring to allow ring flipping to take place (Figure 2).
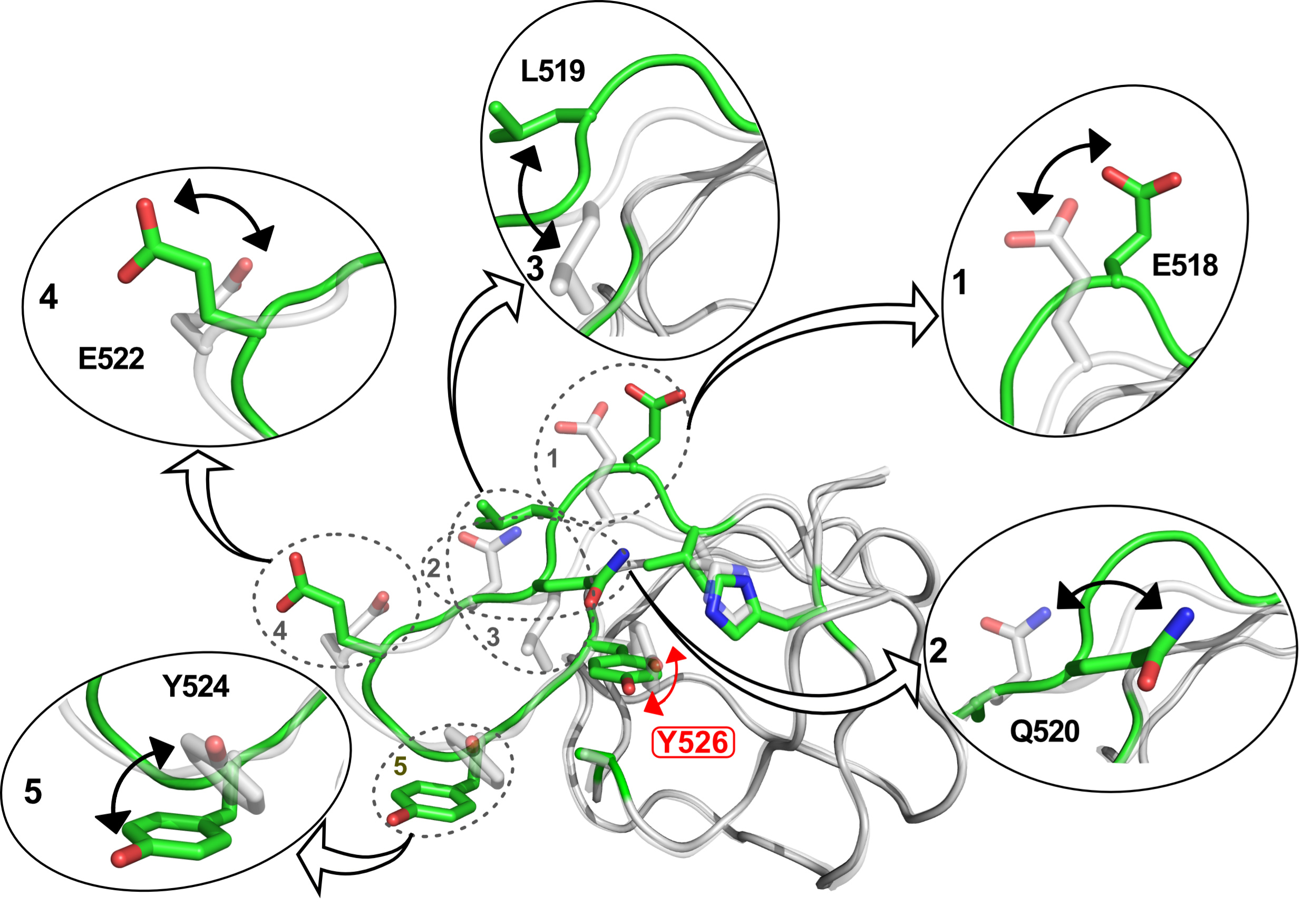
Fig. 1: Illustration of the structural changes associated with rotation of the aromatic ring of the core tyrosine residue (Y526) in the SH3 domain of the MAPK scaffold protein JIP1. The structural changes were captured by individual high-resolution crystal structures of the protein for different rotameric states of the side chain of Y526.
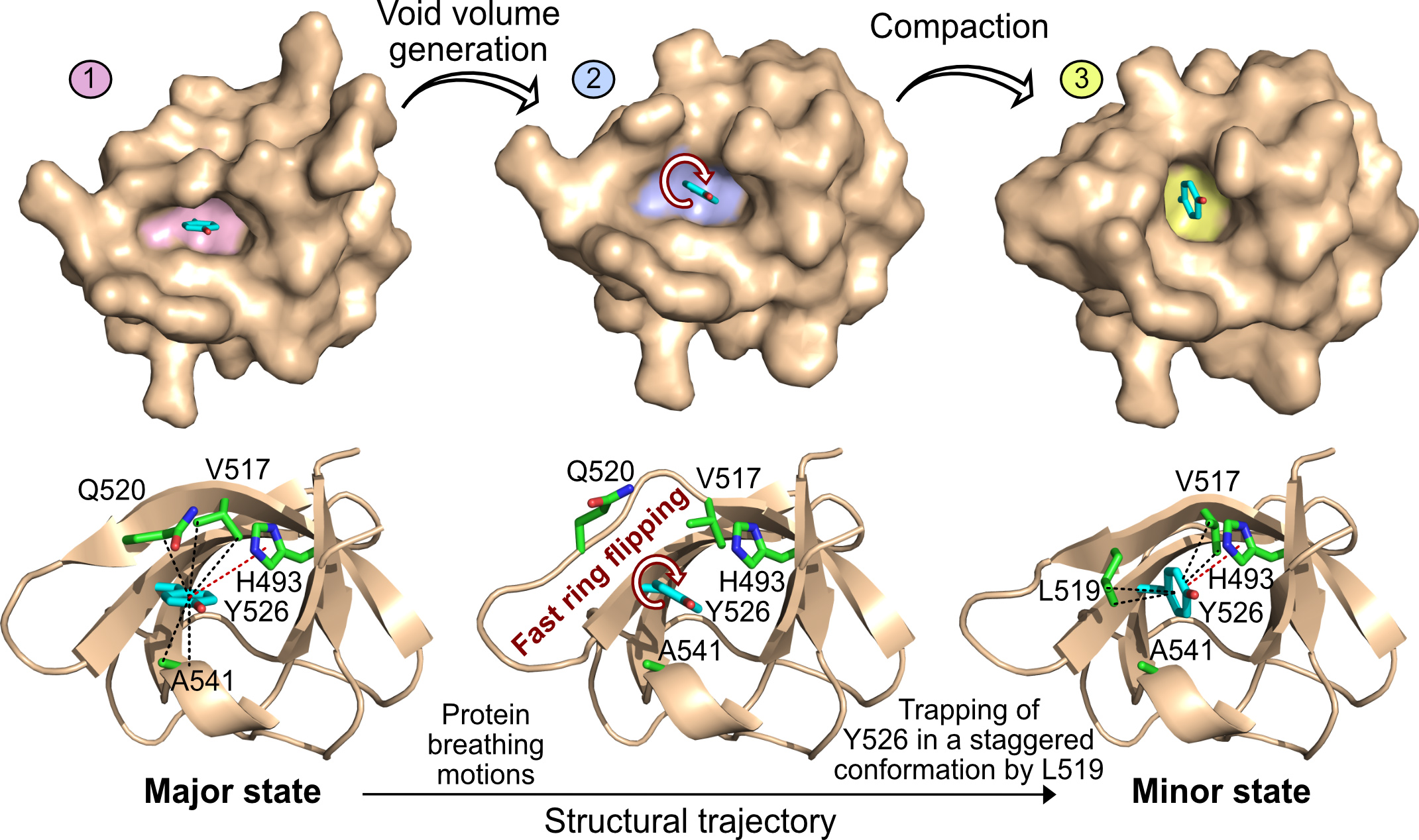
Fig. 2: Top: Surface representation of the SH3 domain of JIP1 in three different rotameric states of Y526 corresponding to the major state, an intermediate state on the structural trajectory and the minor state. The Y526 pocket is highlighted in pink (major), blue (intermediate) and yellow (minor). These rearrangements generate a void volume around Y526, thereby lowering the transition-state energy of ring flipping and allowing the flipping of Y526. Bottom: Illustration of the protein breathing motions along the structural trajectory from the major to the minor state. A void volume is created around Y526, which allows fast ring flipping to take place. The ring flipping is occasionally interrupted by trapping of Y526 in a staggered conformation through formation of CH–π interactions with L519.
This discovery has implications for both protein design and structure prediction by highlighting how even small changes in the delicate balance of interactions stabilising the core can lead to major changes in the protein structure. In addition, it provides a perspective on how novel biological functions can be acquired during evolution through modifications of the intricate network of interactions in the protein core, such as hydrogen bonds, CH-π and π-π stabilising interactions. Thanks to the ability to “see” what is happening in the core of the structures of moving proteins, we get insights into how very small alterations can be the cause of many diseases.
Principal publication and authors
Visualizing protein breathing motions associated with aromatic ring flipping, L. Mariño Pérez (a), F.S. Ielasi (b), L.M. Bessa (a), D. Maurin (a), J. Kragelj (a), M. Blackledge (a), N. Salvi (a), G. Bouvignies (c), A. Palencia (b), M.R. Jensen (a), Nature 602, 695-700 (2022); https://doi.org/10.1038/s41586-022-04417-6
(a) Institut de Biologie Structurale, Université Grenoble Alpes, CEA, CNRS, Grenoble (France)
(b) Structural Biology of Novel Targets in Human Diseases, Institute for Advanced Biosciences, Université Grenoble Alpes, INSERM, CNRS, Grenoble (France)
(c) École Normale Supérieure (ENS), PSL University, Sorbonne Université, CNRS, Paris (France)



