- Home
- News
- Spotlight on Science
- Crystal structure...
Crystal structure of a novel class of ubiquitin E3 ligase
26-06-2018
The cellular regulation by ubiquitin is better understood thanks to a high-resolution structure of MYCBP2, a novel ubiquitin-ligating enzyme from a new class of RCR E3 ligase. MYCBP2 uses a striking tandem cysteine mechanism and transfers ubiquitin to threonine residues via a chemically-distinct esterification reaction. As MYCBP2 has been implicated in neurological disorders, this structure paves the way for novel therapeutic opportunities.
Protein ubiquitination is a fundamental post-translational modification that regulates a plethora of cellular processes [1]. Ubiquitination involves the covalent attachment of the small protein ubiquitin to substrates thereby regulating their stability, localisation or interactions with other proteins. E3 ligases operate at the end of a multienzyme cascade (E1-E2-E3) and typically conjugate the carboxy terminus of ubiquitin to the side chain amino group of lysine residues forming an amide-linked isopeptide bond. There are hundreds of E3 ligases that are believed to be made up of only three main classes – HECT, RING and RBR [2]. An experiment at beamline ID23-1 led to the structure determination of a fourth class of E3 ligase. The RING-Cys-Relay (RCR) E3 found in the neuronal MYCBP2 protein has a mechanism and substrate specificity that redefines the way we should think about ubiquitination.
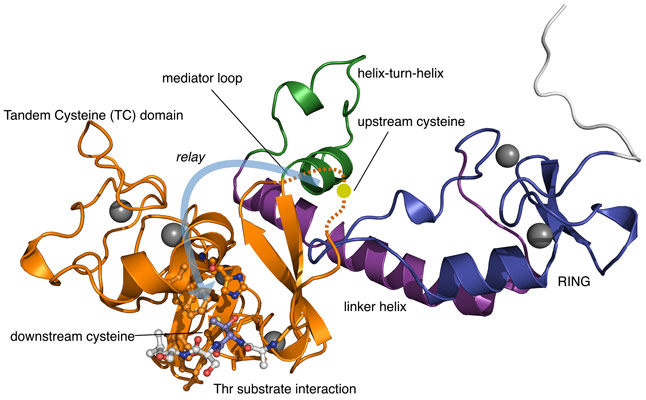
HECT and RBR E3s bind the upstream ubiquitin ‘charged’ E2 enzymes and form a covalent intermediate with ubiquitin through linkage to a single catalytic cysteine residue. The RCR E3 doesn’t have one catalytic cysteine but two! (Figure 1). The upstream cysteine accepts ubiquitin from the upstream E2 and the residue resides within a dynamic region termed the mediator loop. This loop region was found to be disordered within the obtained structural model. It is proposed that the inherent flexibility of this mediator loop, supported by biochemical experiments, allows the RCR E3 to transfer the attached ubiquitin molecule to a downstream catalytic cysteine residue. The two cysteine residues are found in a novel zinc-binding protein fold dubbed the tandem cysteine domain. Such a ubiquitin relay mechanism has never been observed for E3s before but is reminiscent of the protein splicing mechanism of inteins [3].
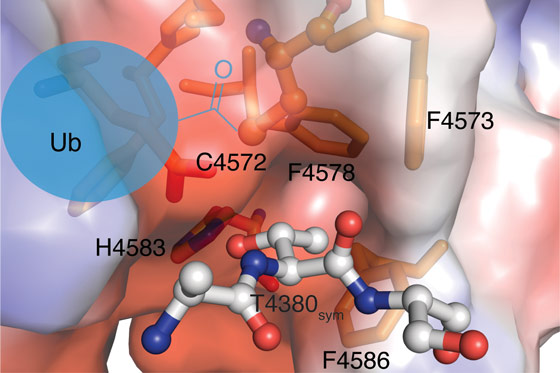
Why the RCR E3 enzyme uses the relay mechanism is not clear but it may explain its surprising substrate specificity. Once the downstream cysteine becomes charged with ubiquitin, it transfers to threonine sides chains thereby forming a chemically distinct ester linkage between substrate and ubiquitin. Insights into the structural basis of this threonine specificity were obtained from a fortuitous packing interaction in the crystal. An N-terminal threonine motif in the crystallisation construct was found to be docked into the active site of a neighbouring molecule centred around the downstream catalytic cysteine (Figure 2). A histidine residue appears to serve as a general base toward the side chain hydroxy group. Furthermore, the methyl group of the threonine side chain docks into a hydrophobic pocket formed by a cluster of three phenylalanine residues. One explanation for the relay mechanism is that these active site requirements for threonine specificity are incompatible with direct transfer of ubiquitin from the E2 enzyme. The downstream cysteine appears to be otherwise quite inaccessible but the mediator loop is optimally tuned to deliver the bulky ubiquitin cargo to it.
This discovery was an interdisciplinary study combining elements of organic chemistry, biochemistry, proteomics and crystallography. Initial evidence that MYCBP2 might be a novel class of E3 ligase was obtained through application of chemical biological tools known as activity-based probes (ABPs) [4]. ABPs covalently label enzymes that demonstrate a specific type of enzymatic activity. When the authors found that their ABPs were labelling MYCBP2 in cell extracts it suggested there might be something particularly interesting about this E3.
As MYCBP2 has a role in driving a neurodegenerative process known as Wallerian degeneration, the structural characterisation of this novel, and seemingly unique, E3 ligase machinery may offer therapeutic opportunity from the treatment of a range of neurological conditions.
Principal publication and authors
Activity-based E3 ligase profiling uncovers an E3 ligase with esterification activity, K.-C. Pao (a), N.T. Wood (a), A. Knebel (a), K. Rafie (b), M. Stanley (b), P.D. Mabbitt (a), R. Sundaramoorthy (b), K. Hofmann (c), D. v. Aalten (b) and S. Virdee (a), Nature, 556, 381–385 (2018); doi: 10.1038/s41586-018-0026-1.
(a) MRC Protein Phosphorylation and Ubiquitylation Unit, University of Dundee, Scotland (UK)
(b) Division of Gene Regulation and Expression, School of Life Sciences, University of Dundee, Scotland (UK)
(c) Institute for Genetics, University of Cologne (Germany)
References
[1] A. Hershko & A. Ciechanover, The ubiquitin system, Annu. Rev. Biochem. 67, 425–479 (1998).
[2] N. Zheng & N. Shabek, Ubiquitin Ligases: Structure, Function, and Regulation. Annu. Rev. Biochem. 86, 129-157 (2017).
[3] K.V. Mills et al., Protein splicing: how inteins escape from precursor proteins. J. Biol. Chem. 289, 14498-505 (2014).
[4] D.S. Hewings et al., Activity-based probes for the ubiquitin conjugation-deconjugation machinery: new chemistries, new tools, and new insights, FEBS J., 284, 1555–1576 (2017).