ESRF’s high-pressure freezing laboratory for macromolecular crystallography
The ESRF’s High-Pressure freezing laboratory for Macromolecular Crystallography (HPMX) is dedicated to investigating interactions between macromolecules and gases in crystallo. Its various high-pressure cells, adapted for different gases, allow the integration of ligands and gas derivatives into macromolecular crystals, complementing diffraction data collection on MX beamlines and providing new insights for functional structural analyses.
Since the early 1990s, bio-crystallographers have used noble gas derivatives to address the phase problem in macromolecular crystallography (MX), subsequently embracing high pressure (helium at 2000 bar) to flash-cool crystals without cryo-protectant. This method continues to be invaluable when cryo-protectant is unavailable to preserve crystalline order or the assembly of macromolecular complexes.
Since 2015, the ESRF has advanced high-pressure (up to 2000 bar) applications in MX to address the needs of its user community. While some macromolecules crystallise naturally with their functional gas, many require pressure for gas integration. By applying high pressure to macromolecular crystals, researchers can map the internal architecture of proteins, including channels, tunnels and cavities, and surface excavations. This facilitates functional investigations into gas–macromolecule interactions, enzyme reaction mechanisms with gaseous substrates, and conformational fluctuations in proteins or ligand-binding processes (Figure 1). Remarkably, less than 1% of all deposited structures in the Protein Data Bank contain gas molecules, significantly underestimating gas-dependent biological systems. This underscores the necessity for further development of high-pressure systems to gain insight into gas–protein interaction mechanisms.
Click image to enlarge
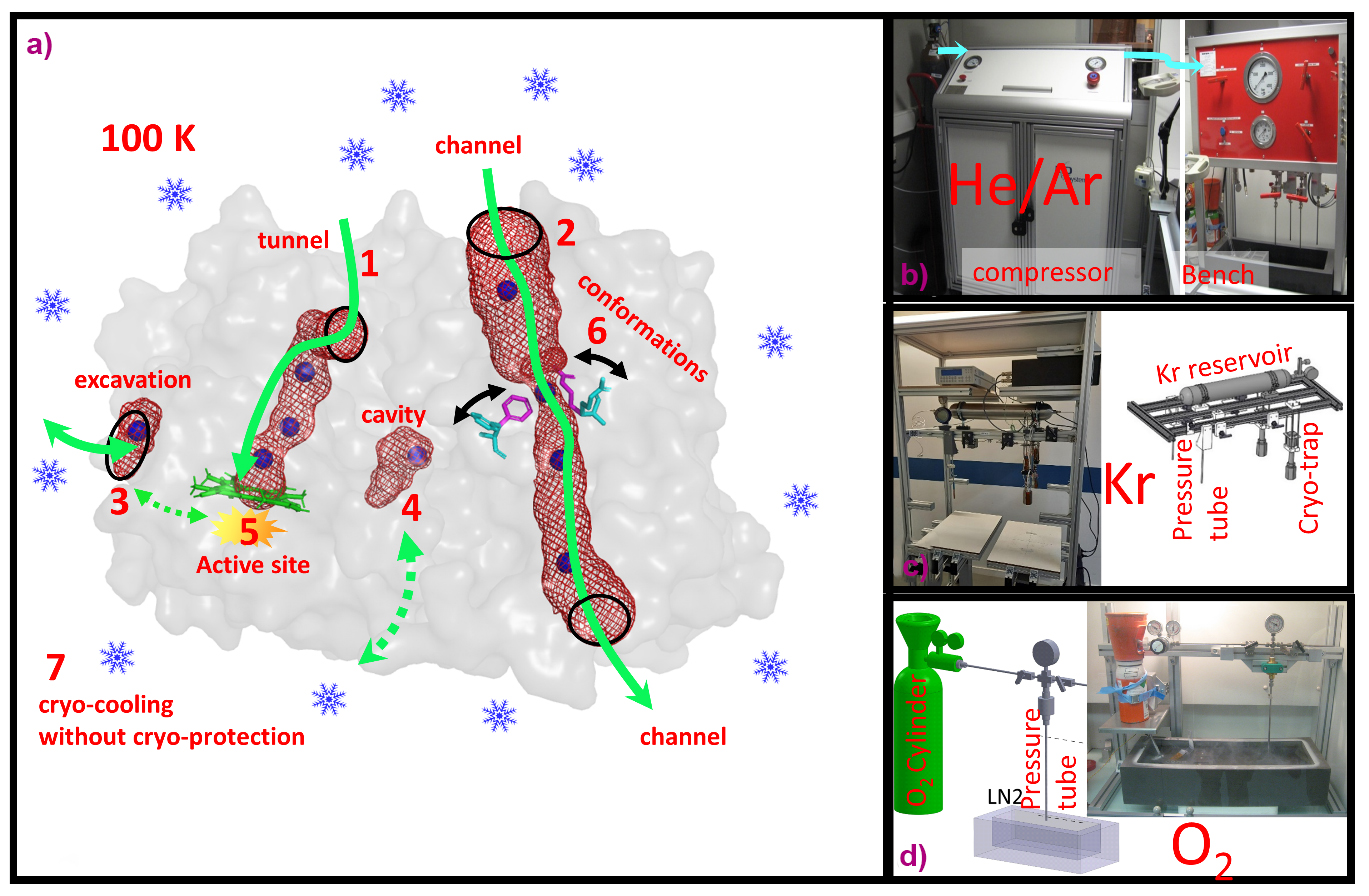
Fig. 1: a) Different study applications of pressurised gases in macromolecular crystallography. 1-4: Mapping of internal and surface architectures of proteins. 5: Reactions of non-inert gases. 6: Exploration of protein conformational fluctuations. 7: Cryo-cooling of crystals without cryo-protectant. b-d) Different pressurisation systems at HPMX for He/Ar, Kr and O2, respectively.
The HPMX currently offers MX users six distinct high-pressure systems, tailored for gases such as helium and argon, krypton, xenon, oxygen, nitrogen and carbon dioxide, and methane. Within these pressure cells, crystals are pressurised to enhance gas–protein interactions, which are then flash-cooled to maintain the pressure-induced state, before depressurisation (also called the ‘soak-and-freeze’ method). Recent examples of high-pressure experiments conducted at the HPMX demonstrate that this innovative method has yielded significant scientific achievements in understanding gas-dependent protein functions.
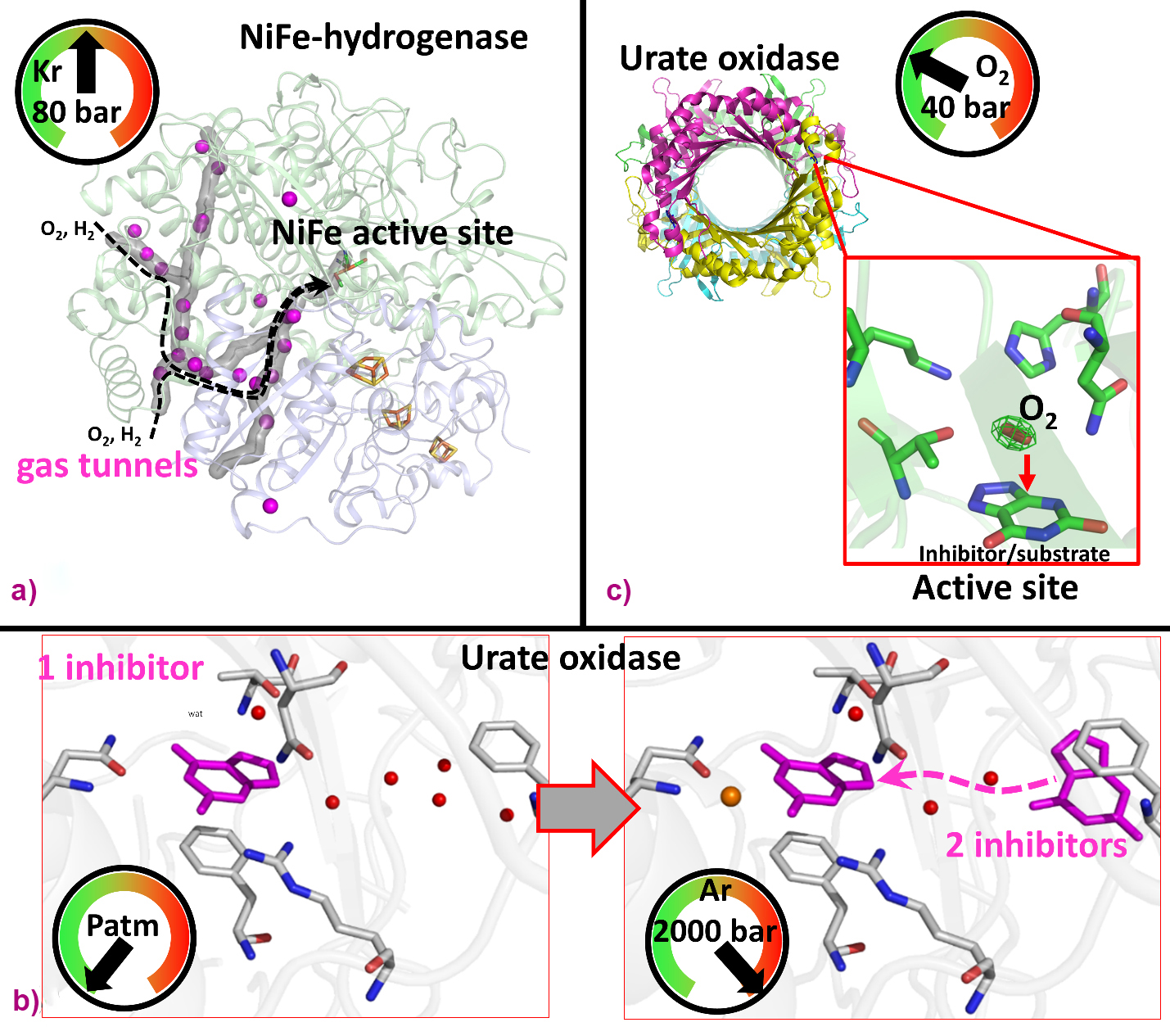
For instance, noble-gas labelling using Xe/Kr enables researchers to map tunnels that facilitate the diffusion of gaseous substrates from the external solvent to the active site of gas-dependent enzymes, such as prototypical hydrogenases catalysing the oxidation of di-hydrogen. Indeed, Kalms et al. used krypton derivatives of hydrogenases to map the hydrophobic channels for H2 (substrate) and possibly O2 (inactivator) diffusion, shedding light on the structural determinants conferring significant oxygen tolerance in selected hydrogenases (Figure 2a) [1].
Click image to enlarge
Fig. 2: Elucidation of protein structural determinants using pressurised gases. a) Mapping of hydrogen diffusion tunnel in an oxygen-tolerant hydrogenase using krypton [1]. b) Analysis of inhibitor binding conformations in urate oxidase using argon [2]. c) Oxygen binding in urate oxidase active site by pressurised O2 [3].
Another study demonstrates that using He/Ar gases at high-pressure (> 1000 bar) can reveal functional structural changes in proteins such as urate oxidase (UOX), which catalyses uric acid degradation (Figure 2b). T. Prangé et al. showed that high pressure stabilises two inhibitors in UOX crystals, instead of one at atmospheric pressure, revealing inhibitor/substrate diffusion via an intermediate position to reach the active site [2].
Additionally, pressurisation using reactant gases (N2, O2, CO2 and CH4) can be used to probe the catalytic mechanisms of gas-employing enzymes, revealing potential intermediates. For example, in a study of UOX crystals pressurised with oxygen, B. Lafumat et al. revealed a specific O2 binding site above an active site inhibitor, well positioned to oxidise the genuine substrate (Figure 2c) [3].
With applications in many fields of research, including fundamental biology, biochemistry, and environmental and medical science, the HPMX has proven to be a valuable ancillary tool for MX users. The provision of pressure cells and their versatile application at the HPMX can be extended to address new scientific questions, tackling challenges faced by modern societies. For example, with fossil resource depletion and rising greenhouse gas levels in the atmosphere, there is an urgent need to study enzymes that transform CO2 and CH4 into valuable chemicals serving as renewable resources. Furthermore, studying the mechanisms of biologically significant macromolecules reliant on light gases, such as hemoproteins binding carbon monoxide and nitric oxide, nitric dioxide reductases and ammonia transporters, could hold implications for health and the treatment of disease [4].
The HPMX is a service of the ESRF’s Structural Biology group, providing the MX community with additional, complementary functionality to the group’s diffraction, scattering and imaging beamlines.
For more information, please visit:
https://www.esrf.fr/home/UsersAndScience/Experiments/MX/About_our_beamlines/HPMX.html
Principal publication and authors
The High-Pressure Freezing Laboratory for Macromolecular Crystallography (HPMX), an ancillary tool for the macromolecular crystallography beamlines at the ESRF, P. Carpentier (a), P. van der Linden (a), C. Mueller-Dieckmann (a), Acta Cryst. D80, 80-92 (2024); https://doi.org/10.1107/S2059798323010707
(a) ESRF
References
[1] J. Kalms et al., Angew. Chem. Int. Ed. 55, 5586-5590 (2016).
[2] T. Prangé et al., Acta Cryst. D78, 162-173 (2022).
[3] B. Lafumat et al., J. Appl. Cryst. 49, 1478-1487 (2016).
[4] I. Melnikov et al., Commun. Biol. 5(1), 360 (2022).