- Home
- News
- General News
- Two transcription...
Two transcription factors together influence your facial appearance
26-01-2024
Researchers led by Stanford University have found that two types of transcription factors involved in embryonic development: TWIST1 and homedomain proteins work together in a coordinated way on the DNA in early development of humans. Their collaboration contributes to the differences in facial appearance among individuals. The scientists solved the structure of the proteins-DNA complex at the ESRF. The results have been published this week in Cell.
Proteins called transcription factors (TFs) control how genes are turned on or off. They have specific DNA patterns they recognize and can bring other helpers to regulate gene activity. In humans, there are large families of TFs like homeodomain (HD) and basic helix-loop-helix (bHLH) proteins that play roles in shaping cellular and positional identities in the developing organism.
To ensure precise gene control, TFs from these large families often collaborate, a process known as cooperative TF binding. This cooperation helps them bind to DNA in a precise way. While there are various ways TFs can cooperate, one less understood method is called "DNA-mediated" cooperativity. Some TFs can work together by binding to neighboring DNA sites without directly interacting with each other. Once bound to DNA in a specific orientation and distance, they make contacts that help stabilise their presence on DNA and are crucial for specific gene regulation.
Although this DNA-guided cooperativity has been observed in lab experiments, its detailed study in living cells is limited to a few examples.
Now an international group of scientists led by Stanford University have studied the protein TWIST1 and its binding to homeodomain TFs and DNA. TWIST1 is transcription factor involved in the regulation of embryonic development. It plays a crucial part in the formation of tissues and organs, including the development of the skeleton and muscles.
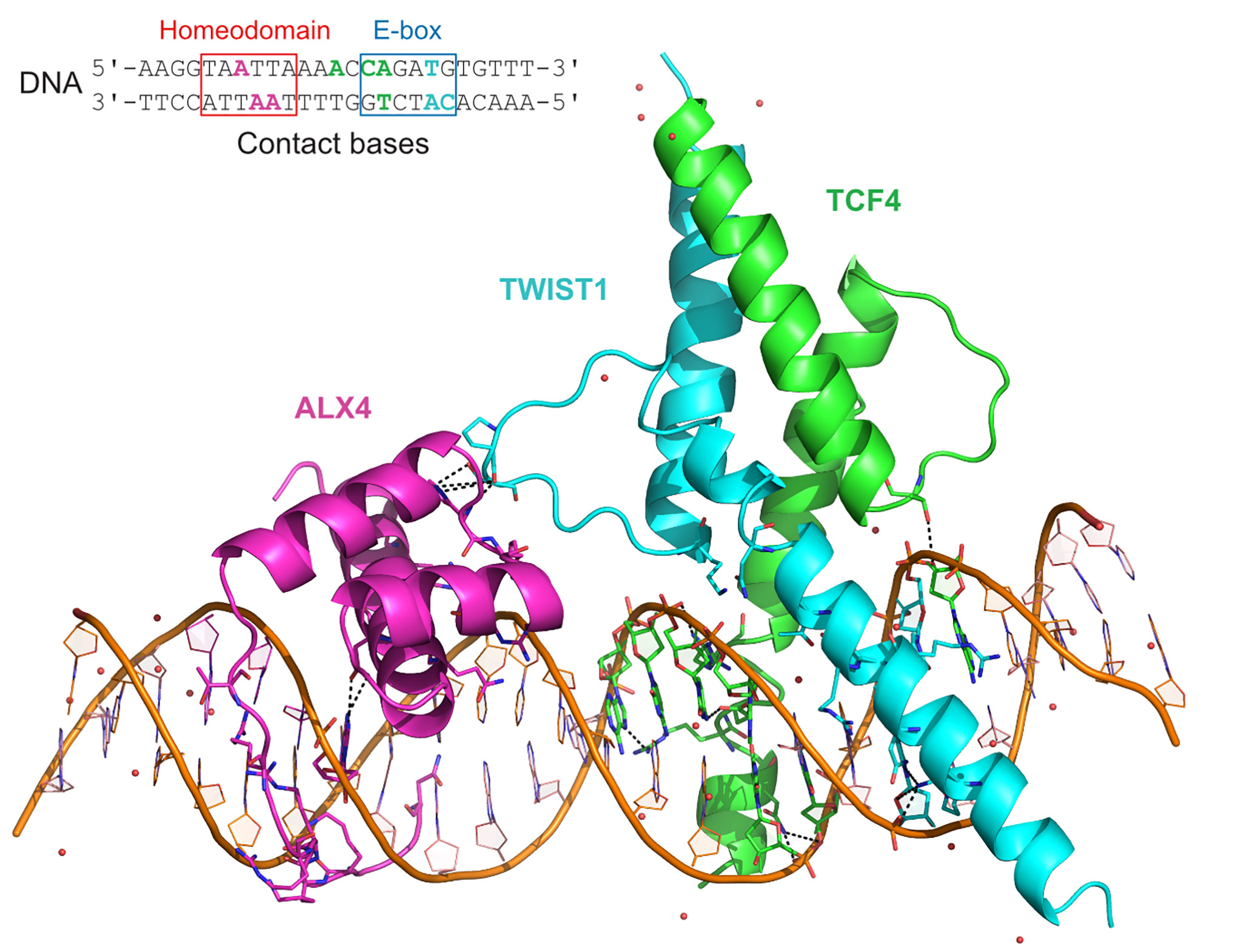
In collaboration with Sasha Popov, scientist at the ESRF and co-author of the paper, the team solved the X-ray crystal structure of the TWIST1, its heterodimerization partner TCF4 and homeodomain factor ALX4 co-bound at a specific DNA sequence at the ESRF’s ID23-1 beamline.
 |
|
3D structure of TWIST1 (aa101–170), TCF4 (aa565–624), and ALX4 (aa210–277) DNA-binding domains bound to the Coordinator DNA sequence. DNA bases recognized by the TFs are highlighted: cyan for TWIST1, green for TCF4, and magenta for ALX4. Credits: Kim, S. et al, Cell, 22 January 2024. |
The researchers found that when TWIST1 and homeodomain TFs ALX4 team up, they enhance the ability of certain regions in the DNA to be accessed, resulting in gene activation. They also discovered that the connections between TWIST1 and homeodomain TFs are not very strong (weak), but they are stabilised by the DNA. This weak interaction helps in choosing specific partners to work together in a selective manner and therefore, leads to shared control over the process of transcription (reading and copying genetic information) in specialized embryonic cell types that give rise to the face and limb. Overall, their findings demonstrate that TWIST1 performs distinct cellular and organismal functions, with a DNA-mediated cooperativity with HD TFs potentially enabling functions that are associated with facial features.
“This research shows how cooperativity guided by the DNA can be highly selective among members of large TF families, in turn resulting in exquisite regulatory specificity that ultimately shapes facial morphology and evolution”, explains Joanna Wysocka, leading scientist of the study and researcher at Stanford University.
Reference:
Text by Montserrat Capellas Espuny



