- Home
- News
- Spotlight on Science
- Engineering a new...
Engineering a new enzyme to metabolise carbon
04-06-2021
Structural data collected at beamline ID23-1 has enabled the directed evolution of a novel, efficient enzyme for carbon assimilation. The evolved enzyme provides an important step towards a circular carbon bioeconomy.
CO2 is a major industrial waste gas and a key driver of global warming. The circular carbon economy envisions the re-purposing of CO2 emissions to make value-added products such as fuels, pharmaceuticals, or plastics. Conversion of CO2 is a difficult reaction and most chemical processes require harsh reaction conditions and lead to a mixture of products. Catalysts lower the activation energy and make the overall process more sustainable and specific. Developing catalysts that can efficiently turn CO2 into multicarbon compounds is therefore a key challenge. In contrast to the established inefficient processes, enzymes operate under benign conditions and typically only generate one product; thus they harbour great potential for carbon capture.
An alternative to direct fixation is to first reduce CO2 into one-carbon (C1) compounds, particularly methanol and formic acid. These compounds can be stored much more conveniently than CO2 and can also be used as feedstock in microbial fermentation. Thus, methanol and formic acid serve as ideal mediators between chemical reduction and biological fixation [1,2].
Additionally, the use of these reduced C1 carriers opens up the possibility of the direct condensation of two C1 units, which is not possible with CO2. Even though C1-C1 condensation reactions are arguably the most elegant solution for carbon fixation, they are rare in nature and only performed by highly complex enzymes. Looking for simpler enzymes, a team of researchers discovered that the low-complexity enzyme MeOXC, which usually breaks a C-C bond, has a slight side-activity towards the condensation of two C1 units.
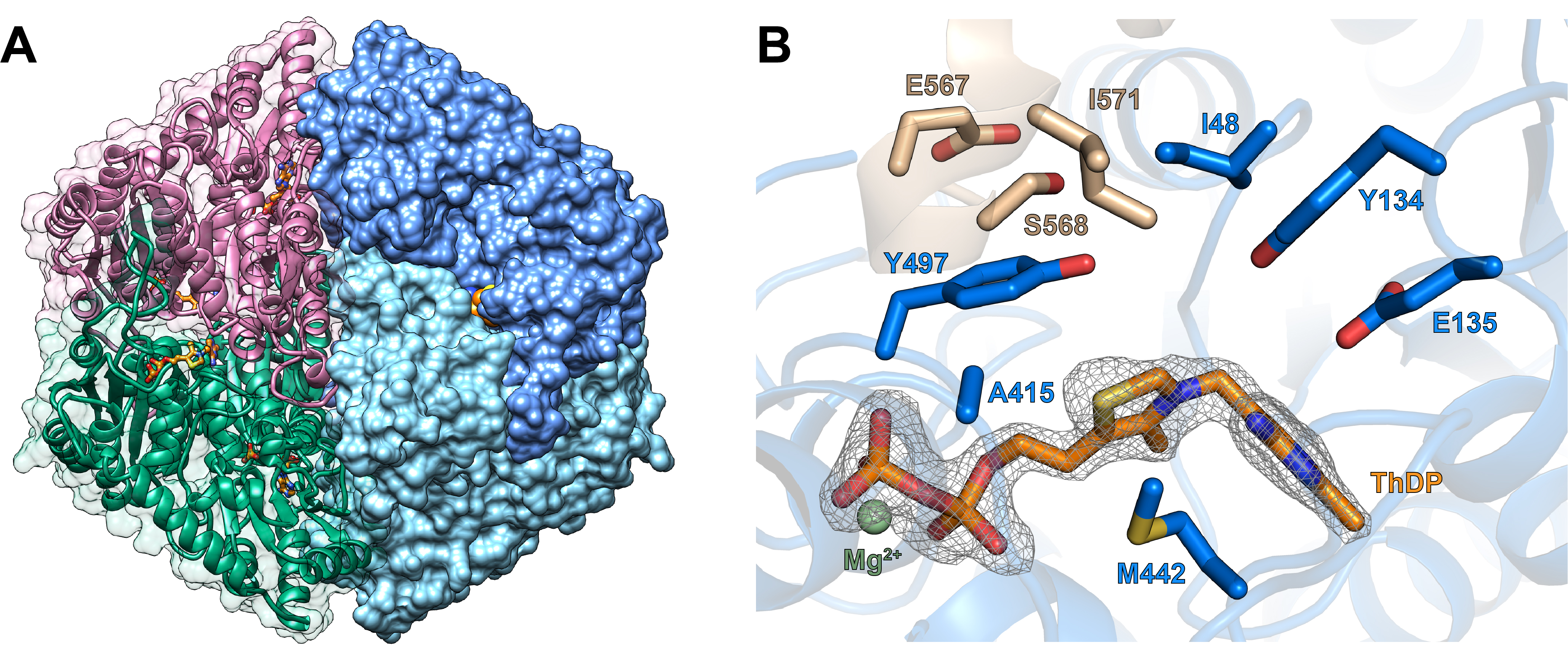
Fig. 1: a) Overview of the overall structure of MeOXC. The cofactor thiamine diphosphate (ThDP) is shown in orange. On the left, a cartoon depiction portraits the folding pattern of two subunits. On the right, a surface representation of the remaining subunits is shown. b) Zoomed-in view of the active site of MeOXC. The ThDP cofactor is shown in orange, and residues within 10 Å of it are depicted as sticks. Blue residues were visible in the crystal structure of MeOXC, while the taupe residues were modelled based on the structure of OXC from Oxalobacter formigenes (PDB ID 2ji7, [3])
Analysis of the enzyme’s crystal structure at beamline ID23-1 gave insight into the active site (Figure 1) and revealed several active site residues that may influence the enzymatic activity. Based on this insight, MeOXC was subjected to directed evolution (Figure 2), a process that mimics natural evolution and uses the principle of survival of the fittest: The enzyme is subjected to random mutagenesis and only the best performing variants are selected for further optimisation. This process is repeated until no improvement is achieved. After four iterations of this process, a quadruple mutant (MeOXC4) with a 2000-fold increased activity for the C1-C1 condensation reaction was obtained. Interestingly, the evolved enzyme had almost fully lost its original activity.
Subsequent crystal structure analysis of MeOXC4 revealed how the evolved enzyme had learned how to accommodate the non-natural substrates and position them for catalysis. The mutations that occurred during the directed evolution abolished a hydrogen-bond network around a water molecule that is important for the enzyme’s natural function. Thus, structural evaluation was not only critical for the design of the novel enzyme but also for understanding the effects of the mutations on the catalytic activity.
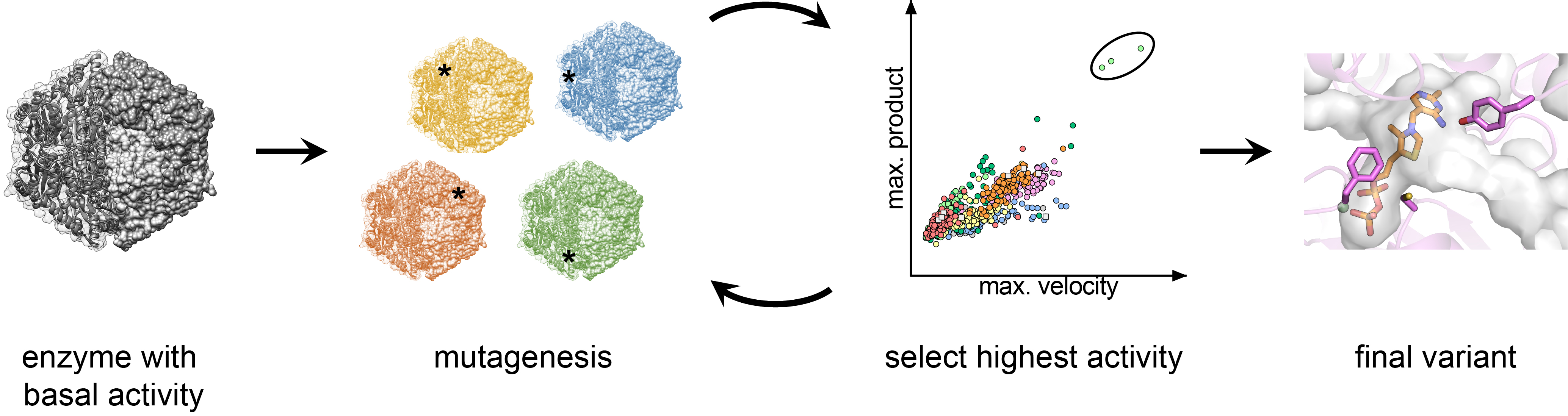
Fig. 2: Enzyme engineering by structure-based directed evolution. After a basal initial activity is detected in the enzyme candidate, its structure is evaluated and targets for mutagenesis are identified based on their proximity to the active site. Variant libraries are created and evaluated in a high-throughput fashion. The best performer is used as template for another round of mutagenesis and screening. Once no further improvement is achieved, the final variant is taken forward to in vivo implementation.
Finally, the engineered enzyme was introduced into Escherichia coli to evaluate its performance in vivo. Similar improvements to the in vitro screen were observed, with every subsequent mutation improving activity of the enzyme in the cellular context. Therefore, the newly designed enzyme is an important step towards creating a microbe that can efficiently turn formate or methanol into valuable products, a process that is essential to realising a circular C1 economy. In a broader context, this work also showcases the potential of protein engineering and synthetic biology in addressing today’s key challenge of carbon emissions.
Principal publication and authors
Engineering a Highly Efficient Carboligase for Synthetic One-Carbon Metabolism, M. Nattermann (a), S. Burgener (a), P. Pfister (a), A. Chou (b), L. Schulz (a), S.H. Lee (b), N. Nicole (a), J. Zarzycki (a), R. Gonzalez (b), T.J. Erb (a,c), ACS Catalysis 11, 5396-5404 (2021); https://doi.org/10.1021/acscatal.1c01237.
(a) Max-Planck-Institute for terrestrial Microbiology, Marburg (Germany)
(b) University of South Florida, Tampa (USA)
(c) LOEWE Center for Synthetic Microbiology, Marburg (Germany)
References
[1] G.A. Olah, Angew. Chem. Int. Ed. Engl. 44, 2636-2639 (2005).
[2] O. Yishai et al., Curr. Opin. Chem. Biol. 35, 1-9 (2016).
[3] C.L. Berthod et al., Structure 15, 853-861 (2007).



