- Home
- News
- Spotlight on Science
- How recruiting a...
How recruiting a novel subunit enabled Rubisco, the carboxylase of photosynthesis, to evolve higher specificity
09-12-2022
Scientists from the Max Planck Institute for Terrestrial Microbiology re-traced the evolution of a novel subunit in the carboxylase Rubisco using X-ray crystallography data from beamline ID23-2. This subunit enabled Rubisco to evolve higher specificities for CO2 over O2.
The appearance of oxygen in the Earth’s atmosphere constituted an important step for the appearance of life as we know it. Organisms, enzymes and metabolisms that arose billions of years ago in anaerobic environments had to adapt to its presence. Among the organisms impacted by the enrichment of oxygen in the atmosphere were those that used Rubisco, the carboxylase of the Calvin cycle, for autotrophic growth. Rubisco reacts both with O2 and CO2, but when the substrate is oxygen it produces an undesired compound that needs to be catabolised through an energy- and carbon-releasing reaction (Figure 1a).
From studying modern forms of Rubisco that evolved in aerobic environments, it was found that one of the ways evolution dealt with Rubisco’s substrate duality was to improve its specificity for CO2 over O2. The molecular determinants of this increased CO2 specificity have remained largely unknown until now, although it was suspected to be linked to the appearance of a novel protein subunit, the so-called small subunit (SSU), in more modern Rubisco species with increased CO2 specificity (Figure 1b). This SSU appeared billions of years ago and is strictly essential in modern Rubisco proteins [1].
To re-trace and understand this event, researchers from the Max Planck Institute for Terrestrial Microbiology and NTU Singapore used a statistical algorithm called “ancestral sequence reconstruction” to recreate forms of Rubisco that could have existed billions of years ago [2], shortly before and shortly after the SSU evolved. The authors demonstrated that the ancestral Rubisco that existed shortly before the emergence of the SSU could not yet interact with it and was not specific. Conversely, the ancestral Rubisco that existed shortly after the SSU’s emergence was already dependent on it and had increased substrate specificity. Importantly, the historical substitutions separating these two ancestral Rubiscos must be responsible for creating the interaction with the SSU and the dependence thereon.
Click to enlarge
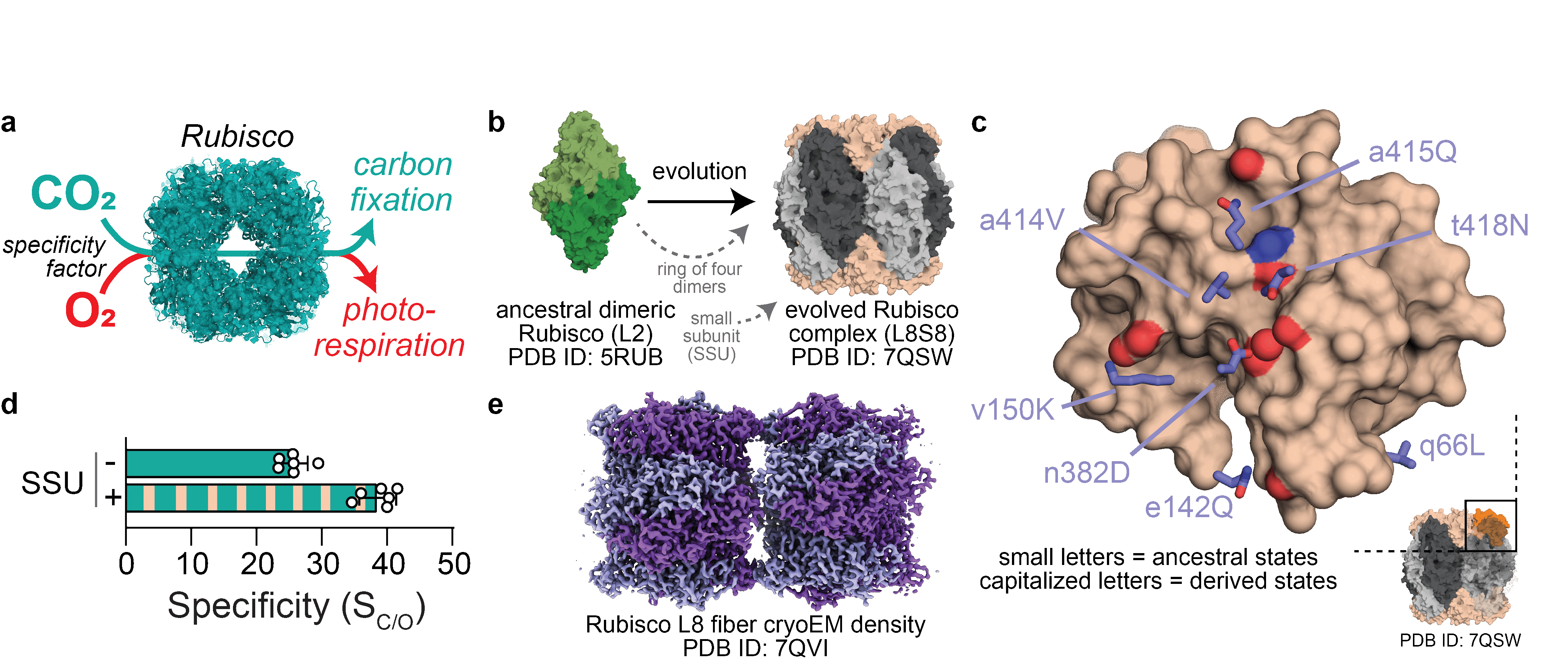
Fig. 1: How Rubisco functions and evolved. a) Ribulose-1,5-bisphosphate carboxylase/oxygenase (Rubisco) can catalyse both a carboxylation and an oxygenation reaction. b) Rubiscos found in aerobic environments evolved from a dimeric assembly of large subunits (L2) to a hexadecameric hetero-complex consisting of eight large and eight small subunits (L8S8). PDB: protein data bank. c) Analysis of Rubisco’s crystal structure yielded important insights into which historical amino acid substitutions occurred in proximity to the small subunit (SSU) binding site, when the SSU first evolved. d) Presence of the SSU increases Rubisco’s specificity for CO2 over O2, enabling better discrimination between the two gases. e) Rubisco evolves to depend on the SSU for solubility and forms insoluble fibres when the SSU is not present.
Analysis of the ancestral Rubisco’s crystal structures at beamline ID23-2 yielded important insights into where in the protein the historical substitutions occurred and which residues were important for the Rubisco-SSU interaction (Figure 1c). Based on these insights, an “intermediate” Rubisco was created that was able to bind the SSU but did not yet depend on it. Using these variants, it was shown that immediately upon its recruitment, the SSU improved Rubisco’s catalysis and opened evolutionary paths that led to increased CO2 specificity (Figure 1d). Notably, the SSU alone was not the cause of drastic changes in specificity. Instead, the acquired domain modulated the effects of further substitutions. Residues without effect in isolation are suddenly influential in SSU presence. The SSU seemingly teleports Rubisco to a new functional sequence space.
A similar effect was responsible for SSU addiction. Accumulation of substitutions that were harmless in the presence of the SSU but deleterious in its absence caused a dependence on it [3]. Rubiscos that accumulated such substitutions formed supramolecular fibrils without the SSU (Figure 1e). These substitutions cause assembly into insoluble fibrils but do not improve Rubisco’s kinetic parameters in any way. Substitutions that made this interaction essential for Rubisco were therefore non-adaptive and accumulated purely because they incurred no negative costs.
These findings suggest that natural selection can fix protein-protein interactions for their beneficial effects and that such interactions can subsequently become essential via non-adaptive evolution. Furthermore, the studied ancestral Rubiscos show that high-specificity arose early in Rubisco’s history, when limited O2 was present [4]. This illuminates a key event in the history of life on Earth. It also reveals how specificity is encoded in Rubisco, which can inform engineering ventures. Instead of being determined solely by changes in the large subunit, the recruitment of a novel interaction partner opened the way for evolution to improve Rubisco’s specificity.
Small accessory subunits can be powerful modulators for the evolution of proteins. This can help access functions that are otherwise not accessible but can equally lead to non-adaptive dependencies that evolve through a shifted accessible sequence space.
Principal publication and authors
Evolution of increased complexity and specificity at the dawn of form I Rubiscos, L. Schulz (a), Z. Guo (b), J. Zarzycki (a), W. Steinchen (c,d), J.M. Schuller (c,d), T. Heimerl (c,d), S. Prinz (e), O. Mueller-Cajar (b), T.J. Erb (a,c), G.K.A. Hochberg (a,c,d), Science 378, 155-160 (2022); https://doi.org/10.1126/science.abq1416
(a) Max Planck Institute for Terrestrial Microbiology, Marburg (Germany)
(b) Nanyang Technological University Singapore (Singapore)
(c) Center for Synthetic Microbiology (SYNMIKRO), Marburg (Germany)
(d) Philipps University Marburg (Germany)
(e) Max Planck Institute for Biophysics, Frankfurt (Germany)
References
[1] T.J. Erb et al., Curr. Opin. Biotech. 49, 100-107 (2018).
[2] G.K.A Hochberg et al., Ann. Rev. Biophys. 46, 247 (2017).
[3] G.K.A. Hochberg et al., Nature 588 (7838), 503-508 (2020).
[4] P.M. Shih et al., Nat. Commun. 7, 10382 (2016).
| About the beamline: ID23-2 |
|
ID23-2 is a fixed energy beamline dedicated to macromolecular crystallography (MX). It is particularly specialised to perform experiments with very small protein crystals and for serial crystallography. ID23-2 delivers a flux of about 1x1013 ph/s in a microfocused beam of ~5x5 µm. Since the recent upgrade, the beam is vertically focused by CRLs and horizontally focused by a mirror in half-Kirkpatrick-Baez (KB) geometry. The beamline is equipped with a MD3-UP microdiffractometer with a vertical spindle that can perform very fast mesh scans and is compatible with in-situ data collection for low-profile crystallisation plates. A FlexHCD sample changer robot has been installed that can take up to 23 Unipucks. The detector is a PILATUS3 X 2M with a maximum frame rate of 250 Hz for the full surface and 500 Hz when using exclusively the central modules of the detector. |